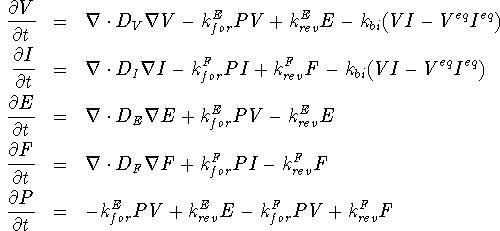
This example demonstrates a strongly coupled highly reactive model for phosphorus diffusion [4].
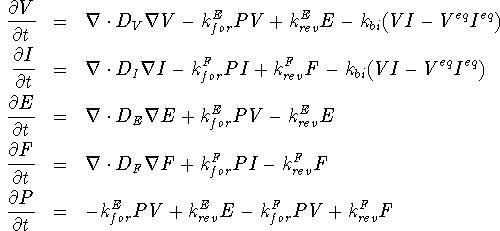

The following script shows the model definition.
# This is the Mulvaney & Richardson 5-species kinetic model for Phosphorus
# diffusion.
# set up some TCL variables to alias field names
set V VacancyInSilicon
set I InterstitialInSilicon
set P PhosphorusInSilicon
set E PVpairInSilicon
set F PIpairInSilicon
# parameters used by Mulvaney & Richardson
set Dv 0.6e-8
set Di 6e-8
set De 6e-12
set Df 1.2e-11
set Kfe 6e-13
set Kre 6e2
set Kbi 6e-9
set Kff 6e-13
set Krf 7.2e2
set Vstar 1e14
set Istar 1e14
set SurfConc 3.2e20
# Create each equation
set vacancyEquation [equation -parabolic \
-name "Vacancy Equation" -solvesFor $V \
-operators [list \
[operator diffusion -sign 1 -parameters [list Dv $V] ] \
[operator k1fg_k2h -sign -1 -parameters [list $P $V $E Kfe Kre] ] \
[operator bimole -sign -1 -parameters [list $I $V Kbi Istar Vstar] ] ] ]
set interstitialEquation [equation -parabolic \
-name "Interstitial Equation" -solvesFor $I \
-operators [list \
[operator diffusion -sign 1 -parameters [list Di $I] ] \
[operator k1fg_k2h -sign -1 -parameters [list $P $I $F Kff Krf] ] \
[operator bimole -sign -1 -parameters [list $I $V Kbi Istar Vstar] ] ] ]
set pvEquation [equation -parabolic -name "PV-pairs Equation" -solvesFor $E \
-operators [list \
[operator diffusion -sign 1 -parameters [list De $E] ] \
[operator k1fg_k2h -sign 1 -parameters [list $P $V $E Kfe Kre] ] ] ]
set piEquation [equation -parabolic -name "PI-pairs Equation" -solvesFor $F \
-operators [list \
[operator diffusion -sign 1 -parameters [list Df $F] ] \
[operator k1fg_k2h -sign 1 -parameters [list $P $I $F Kff Krf] ] ] ]
set pEquation [equation -parabolic -name "Phos Equation" -solvesFor $P \
-operators [list \
[operator k1fg_k2h -sign -1 -parameters [list $P $V $E Kfe Kre] ] \
[operator k1fg_k2h -sign -1 -parameters [list $P $I $F Kff Krf] ] ]]
The model will be used to simulate a predeposition step. The boundary conditions needed are dirichlet boundary conditions which fix the values of P, I, and V at the exposed surface.
# Combine the equations into a model with Dirichlet BC
set modelDiff [model \
-systems [list \
[systemInRegion -name "Kinetic Phos" -region Silicon -equations [list \
$vacancyEquation \
$interstitialEquation \
$pvEquation \
$piEquation \
$pEquation ] ] ] \
-fixedBC [list \
[list $V ExposedSilicon] \
[list $P ExposedSilicon] \
[list $I ExposedSilicon] ] ]
Once the structure is read, the values in the structure file are discarded, and the fields are initialized with flat profiles.
# Bring in the structure file structure -infile init.str # Overwrite any initial values in the structure file field $V -setValue $Vstar field $I -setValue $Istar field $E -setValue 1.0 field $F -setValue 1.0 field $P -setValue 1.0 # Set the surface concentration. This must be *after* setting the # interior P concentration. field $P -inRegion ExposedSilicon -setValue $SurfConc # Tune the timestepping/nonlinear algorithms (all of this could be # configured in the call to model above. # Use a relative nonlinear tolerance of $modelDiff -relNLTol 1.0e-9 # and an initial time step of $modelDiff -deltaT 1.0e-4 # never giving up in the nonlinear solve $modelDiff -maxNLSteps 0 # taking an extra step for good measure $modelDiff -extraNLSteps 1 # tune the SLES linear solver to use ILU preconditioning and a # bi-conjugate gradient squared accelerator $modelDiff -slesPreconditioning ilu -slesAccelerator bcgs # Stop after 10 (minutes, in this case) $modelDiff -stopTime 10 # Now, try to solve it. $modelDiff -transientSolve # Spew out the updated results structure -outfile final.str